Researchers looking for a test to predict whether someone has a potentially deadly condition called sepsis have made a surprise finding – they can predict who will die from it.
Their test accurately picked out who would develop severe sepsis and die, versus those who had fairly innocuous infections and lived.
They hope they can develop it into a tool that will help doctors decide who needs immediate hospitalization and intensive treatment, versus those who can tough it out. Eventually, they hope they may be able to predict who might be susceptible to infections of all kinds.
“The test is looking for a signature that will guide a physician – is this patient going to have a catastrophic illness or is he going to have a mild infection?” said Dr. Stephen Kingsmore, president and chief executive of the National Center for Genome Resources in Santa Fe, N.M., who led the team.
Sepsis is the 10th leading cause of death in the United States. It’s still a mystery to doctors. At first defined as a blood infection, it’s now more widely understood as a devastating reponse to infections and injuries of many different kinds. Patients can go from being perfectly well to being on the verge of death. They can lose limbs, and die within hours.
The number of people treated in the hospital for sepsis in the U.S. has more than doubled in recent years, to about 1.6 million a year, according to a 2011 report from the Agency for Healthcare Research and Quality. There’s not a good test for sepsis, so it’s not clear how many people who develop sepsis die of it.
Kingsmore’s team started looking for that answer. They took a unique approach, running a test for just about every protein and metabolite in human blood. That’s tens of thousands of compounds, in a study akin to mapping the entire human genome, says Kingsmore.
“You measure everything and you see what falls out. That is what is kind of cool about this study,” Kingsmore said in a telephone interview.
They tested 1,152 people who developed sepsis from 2005 to 2009 at Henry Ford Hospital in Detroit, and Duke University Hospital and Durham Veterans Affairs Medical Center in North Carolina. The patients had their blood sampled when they first came in to be treated, and then later on, also.
“What we found in this study was really surprising to us,” Kingsmore told NBC News. “The study was designed to identify a signature that would just tell us if it was sepsis or not because it so difficult for a doctor to identify sepsis. Much to our surprise, as we got into the study and identified the results, we found that we have these really big differences between these patients.”
The patients who developed severe sepsis or died had different levels of many different blood chemicals, the team reports in the journal Science Translational Medicine.
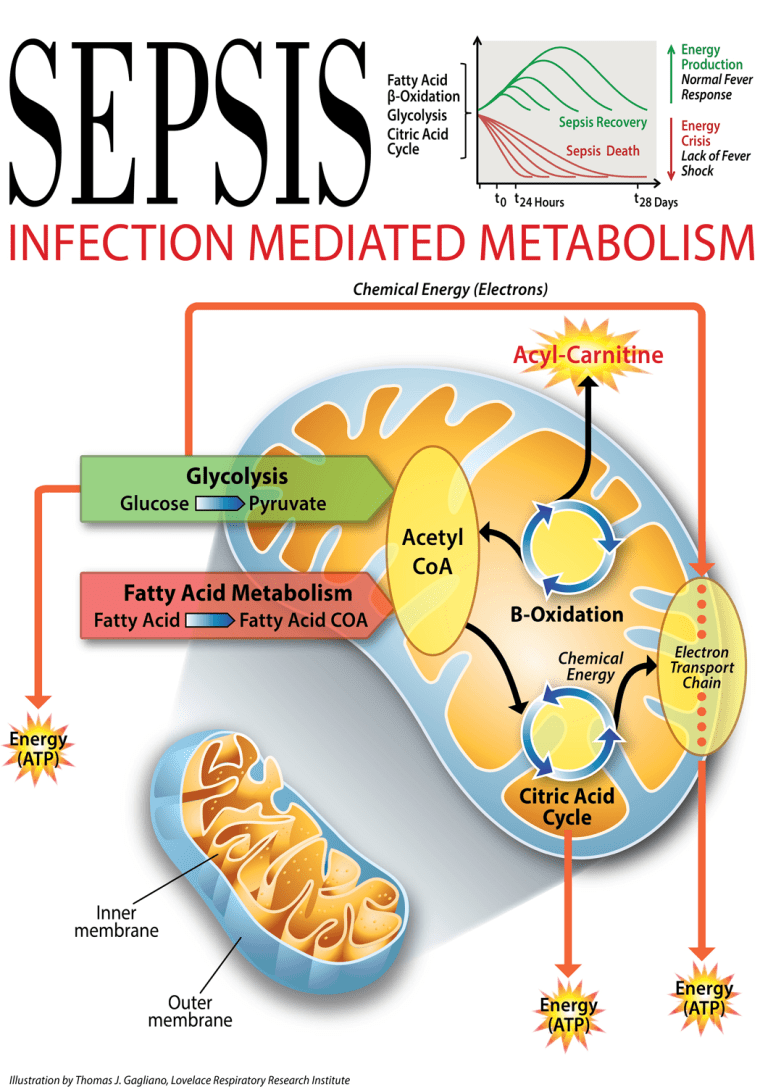
“There were hundreds or differences,” Kingsmore said. Surprisingly, it didn’t matter which bacteria was causing the infection – the signature was similar. There was a clear pattern, and it involved how cells use energy.
“They are measures of the body’s ability to metabolize, to generate energy,” Kingsmore said. “One group had a robust body response to serious infection. They generate a lot of energy and they have a fever. In ways we don’t understand, that contributes to their survival.”
But many of the patients did not do well. Their blood tests, taken when they first showed up at the hospital, showed there were already differences. “In people with bad outcomes, they don’t have that response," Kingsmore said. "They have a kind of partial response. They mobilize fatty acids, but their cells are not able to harvest that energy.”
Much more testing is required to find out just what is happening in these patients, noted Kingsmore, who also heads the pediatric genomic medicine program at Children’s Mercy Hospital in Kansas City.
But he believes it has great potential. “This signature is going to be good for a broad spectrum of infectious diseases. That would include the flu and other severe viral infections as well as the common bacterial infections and even infections with fungi,” he said.
It may even be a signal for patients who will die of trauma, perhaps after a car accident, he said.
The team tested their findings over and over again to be sure, said Kingsmore. The work lasted eight years.
“We talk about survival of the fittest and here have defined the fittest in a new way, a really surprising way,” he said. “The people don’t have the type of robust response that you need to survive an infection. The hope is we can reverse that if we identify the patients early,” he added.
“The goal is not to tell a patient you are going to die.”
That’s a new challenge, because there are not any good drugs for sepsis. There’s a device that identifies the most likely bacteria to cause sepsis, but no highly effective drugs. Patients get intravenous antibiotics and fluids, as well as careful monitoring, but can still die.